GATU Quick Start
What is it? Genome Annotation Transfer Utility (GATU) annotates a genome based on a very closely related reference genome. The proteins/mature peptides of the reference genome are BLASTed against the genome to be annotated in order to find the genes/mature peptides in the genome to be annotated.
What can I do with it? GATU’s main purpose is to allow you to quickly and easily find which regions in an unannotated genome match that of a fully annotated reference genome, and then create a Genbank file of your results.
How do I get started? GATU can be accessed for free at www.virology.ca. From the VBRC Tools drop-down on the left side of the page, select GATU, then click the Launch Program button. You can also access GATU directly from here. GATU will be downloaded to your computer and Java Web Start will automatically start the program.
Note: A warning message that appears which requires users to authenticate GATU. This allows GATU to save save files to your computer.
What next? The main GATU window and its functions are shown below.
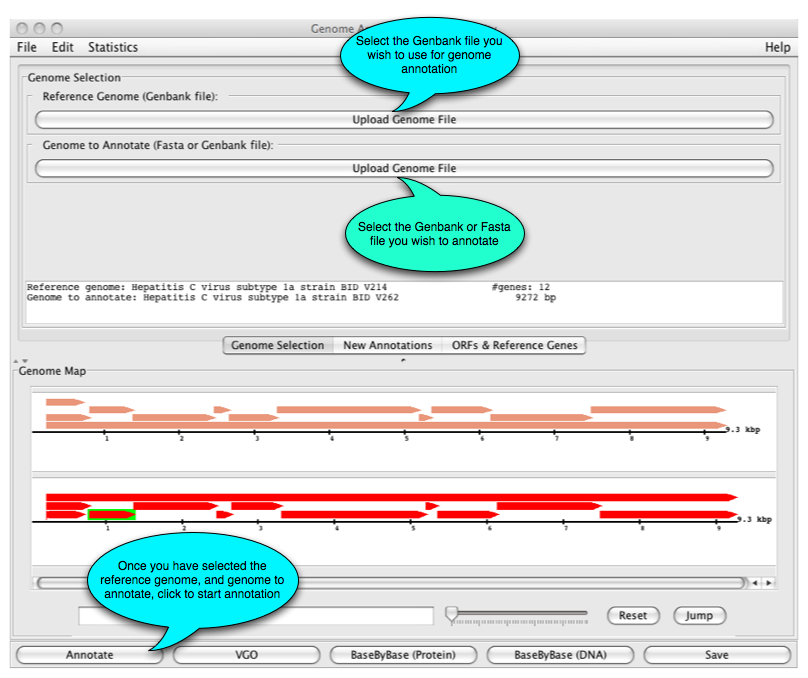
Once you have made your selection, the following display appears:
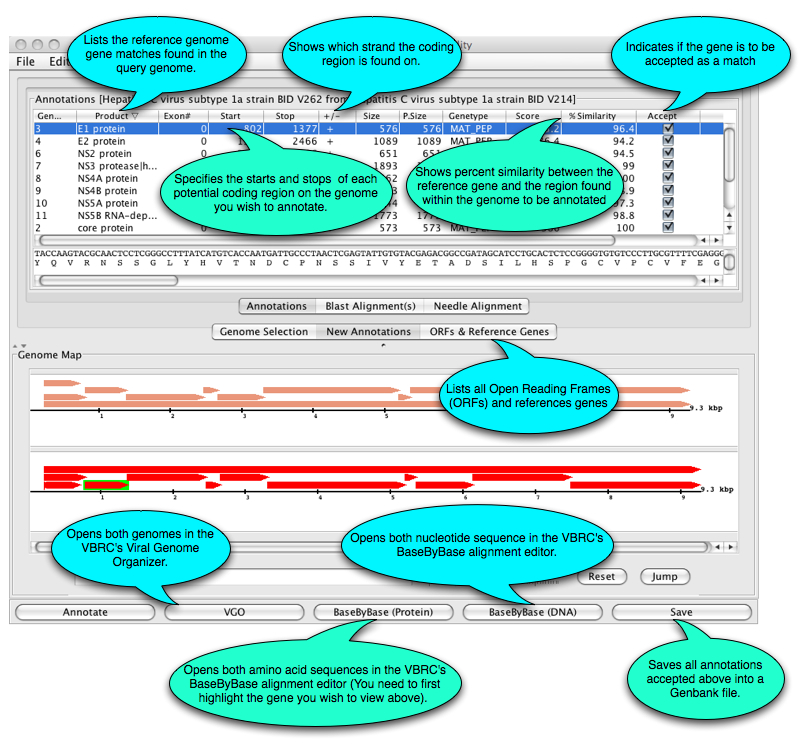
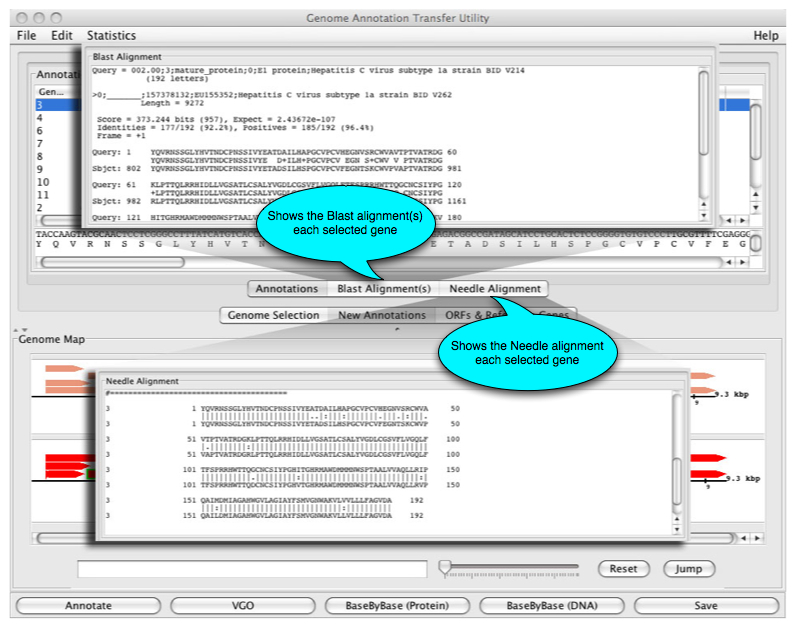
Users can also view the Blast and Needle aligments performed by GATU:
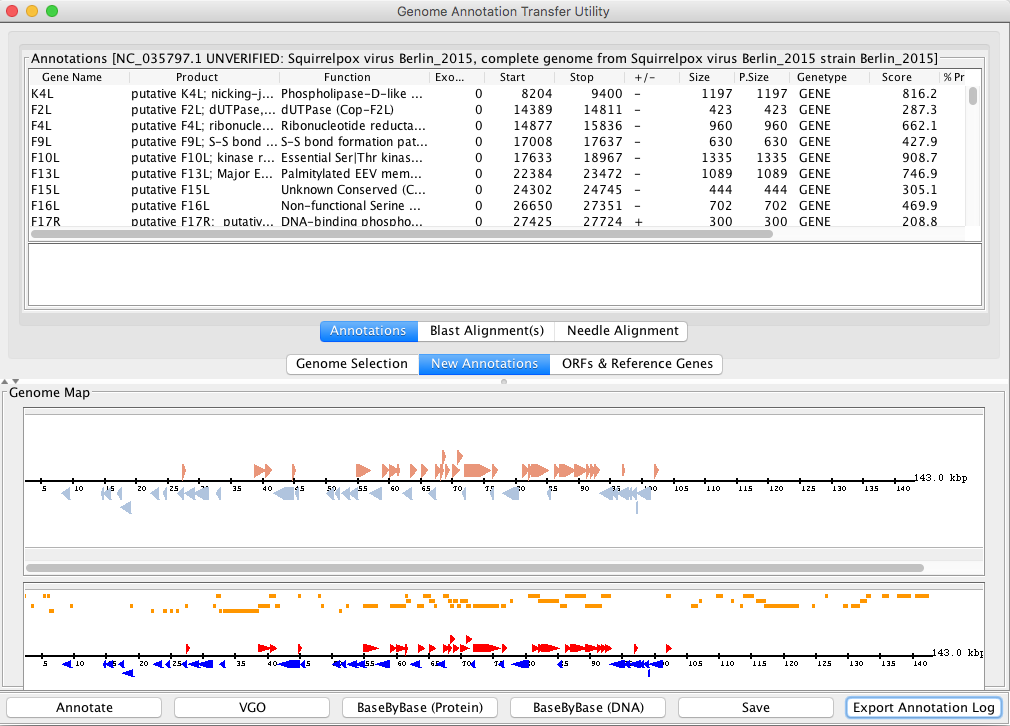
A new button named “Export Annotation Log” has been added. The purpose of this button is to save all accepted and unaccepted annotations and ORFs in a CSV file. In CSV file, the column called “Accepted” has value either TRUE or FALSE to indicate whether annotations and ORFs were accepted or not while exporting the those annotations.