The Genome Data Table
The Genome Data Table Main Window
Within this table, genome names are listed in the first column and corresponding information for each genome is listed in successive columns within the corresponding row.
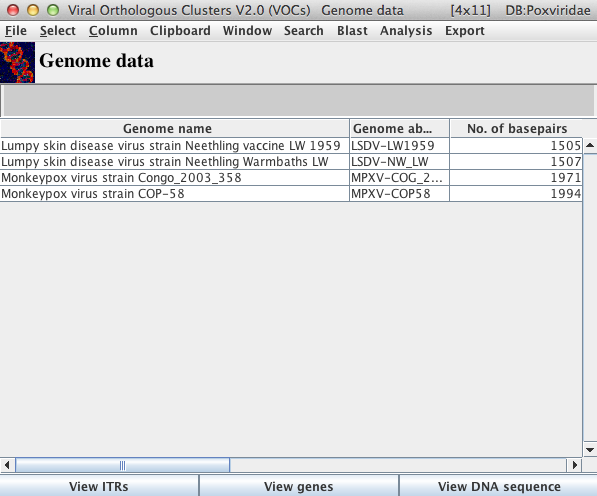
The View ITRs button will open a window for each genome selected to display the corresponding 5′ and 3′ ITR nucleotide sequences (note that this function only applies to Adenoviridae, Asfarviridae, Herpesviridae and Poxviridae).
The View genes button will display one Gene Results Table for all selected genomes.
The View DNA sequence button will display a searchable DNA window with the DNA sequence of the selected genome. This may display either the complete DNA sequence OR the DNA sequence between the first and last codons (inclusive)depending on the GenBank file.
![]()
The functions within the menus “File“, “Select“, “Column” and “Clipboard” are common functions to all tables explained in The Tables of Vocs
The “Search” Menu
From this portion of the menu, you can conduct searches within the selected genomes, for more information on how to use this feature, go to “The “Sequence” Menu” section of The Gene Results Table documentation.
The “Blast” Menu
“BLASTn”
This will allow you to BLAST the sequence (only one genome may be selected at a time) against the NCBI database or any of our databases. The output can be displayed in your choice of HTML or text.
“BLASTn two (partial) sequences”
This is used to BLAST two sequences against each other. The genomes selected may be complete or partial sequences.
The “Analysis” Menu
DNA Grapher will use the entire genome to create a DNA skew graph.
The “Export” Menu
For the selected genomes you may choose to “write (a) GenBank file“, “write (a) FASTA file” and/or “write (an) XML (BSML) file“. If you select to save more than one genome in FASTA or XML (BSML) format, one file with all selected genomes will be created. If you select to save more than one genome to a GenBank file, you will be prompted to save each sequence individually.